The value of the Genome-Phenome Analysis Platform has been recognised by the International Rare Disease Research Consortium (IRDiRC). In April 2018, the tool received the IRDiRC Recognized Resource label, which highlights resources of fundamental importance to the international rare diseases research and development community.
For latest updates, see the platform release notes.
The access to the Genome-Phenome Analysis Platform is provided free of charge for authorised scientists and clinicians around the world. Register here.
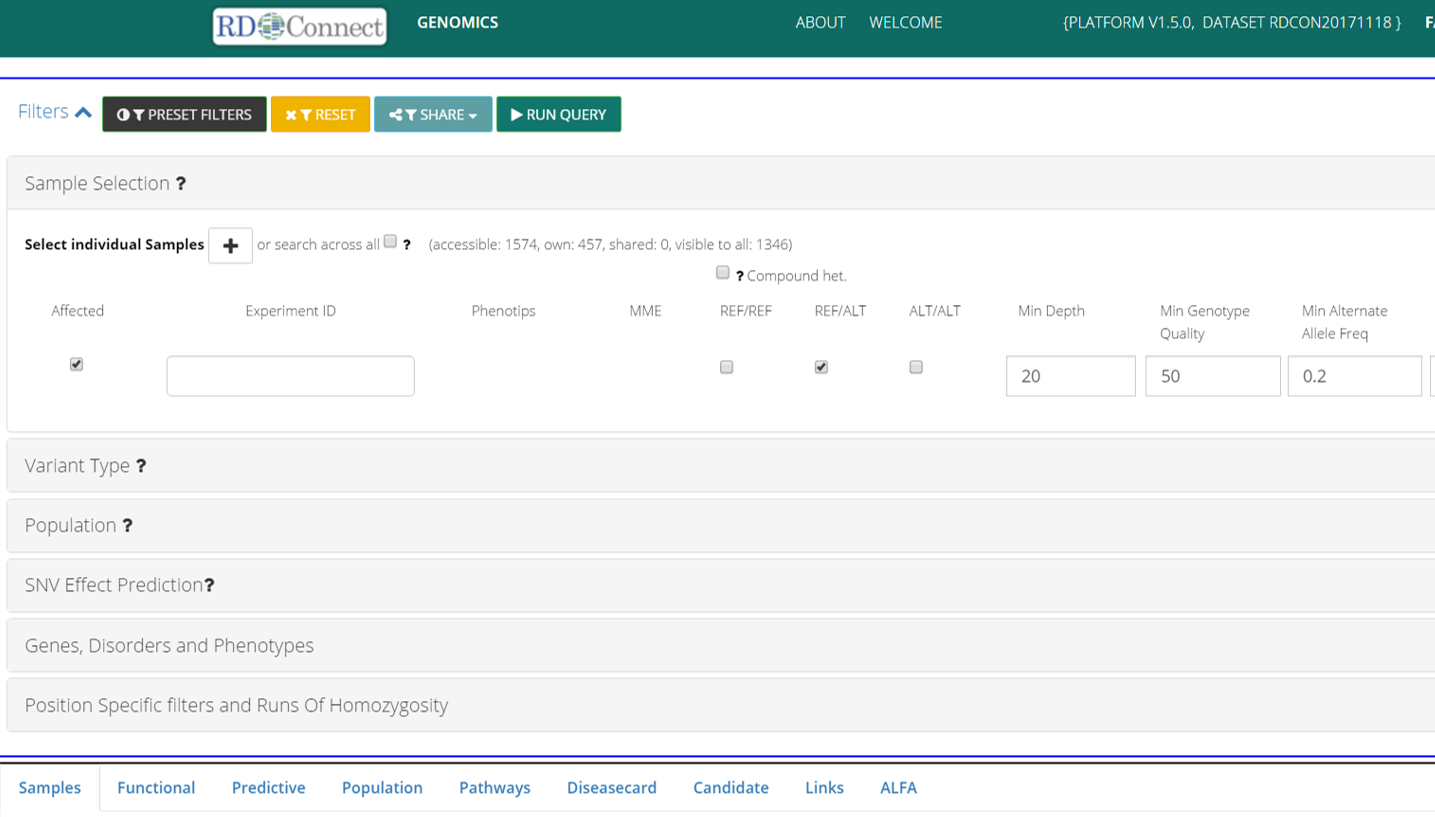
What you need to know about Genome-Phenome Analysis Platform
 The European Genome-phenome Archive (EGA) is a repository for all types of sequencing and genotype experiments, including case control, population, and family studies. It includes genotypes regarding single nucleotide polymorphism (SNPs) and copy number variant (CNVs) from array based methods and genotyping done with re-sequencing methods.
The European Genome-phenome Archive (EGA) is a repository for all types of sequencing and genotype experiments, including case control, population, and family studies. It includes genotypes regarding single nucleotide polymorphism (SNPs) and copy number variant (CNVs) from array based methods and genotyping done with re-sequencing methods.
The EGA serves as archive for publication as well as data on several levels, including the raw data (so they could be re-analysed in the future using other algorithms) and the genotype calls (information about pathogenic genetic variants) provided by the data submitters.
The EGA provides the necessary security required to control access to the data and maintain patient confidentiality. Data can be viewed only by authorised researchers and clinicians. In all cases, data access decisions are made not by the EGA but by an appropriate Data Access Committee.
Who can access the data? The data stored on the Genome-Phenome Analysis Platform can be accessed only by the authorised scientists and clinicians who underwent the registration and verification process.
Embargo period The users can analyse and query their own data as well as data submitted by others. Data become accessible to other authorised users only after a predefined embargo period of six months which allows researchers priority to access to their own data.
Patient security and confidentiality To protect patient privacy, data of individual patients are linked to the unique RD-Connect IDs. Patients’ identity data are not stored on the Platform and cannot be accessed by Platform users. Only the researcher, who submitted the data, has the key to identify the IDs corresponding to his/her data.
Still have questions? Check the GPAP FAQ:
The Platform has been developed at the Centro Nacional de Análisis Genómico (CNAG) in Barcelona, Spain.