The RD-Connect data linkage team focuses on the technical aspect of linking different types of data between institutions across countries. This requires adopting the FAIR principles, meaning the data stored in databases should be Findable, Accessible, Interoperable and Reusable to allow data exchange. FAIR data standards help to harmonise and link up data from different sources, including: genomics archives, health records, patient registries and biobanks.
Rare disease patient registries contain important information to advance the knowledge of diagnosis, disease progression, and treatment, and it is therefore in the patients interest that this data is optimal for research. Making data FAIR for research is especially crucial for rare diseases as they often are sensitive, sparse, highly distributed and heterogeneous.
FAIRification of a rare disease patient registry means that the registry data and metadata are made machine-readable, that the metadata clearly describes how the the data can be accessed and reproduced, and that the metadata can be found by machines. This requires complete understanding of the registry data including:
- the data items/elements;
- how it was created;
- the accessibility restrictions.
Data FAIRifier
FAIRification requires tools, such as Data FAIRifier developed by DTL, including RD-Connect partners.
How to make data FAIR
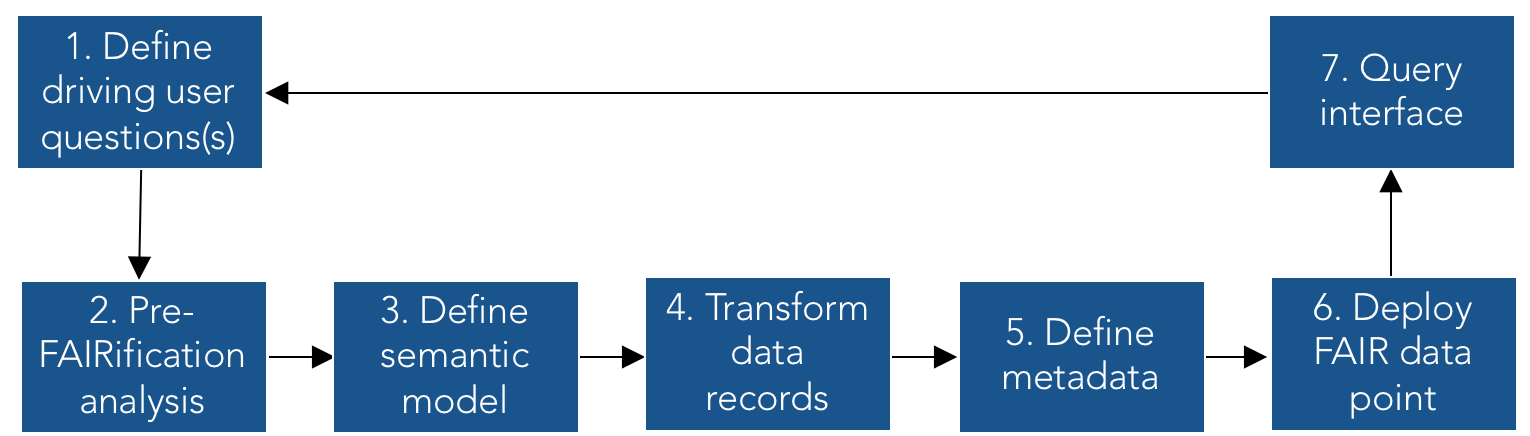
All rare disease databases, such as patient registries, are encouraged to make their data FAIR, which involves 7 steps described below. The process will be slightly different in each individual case, so if you want to FAIRify your database, we recommend that you consult a FAIR data expert first.
Machine-readability
Data and metadata are made interoperable by making them machine-readable. The degree of interoperability is greatly impacted by the choice of ontologies by which data are described. Human Phenotype Ontology and Orphanet Rare Disease Ontology are already IRDiRC Recognized Resources and are being applied by researcher when describing phenotypic data and rare diseases. Data and metadata concepts and relations should be described by globally unique and persistent identifiers. Thus the original data (which could be tabular) is converted to an ontology-grounded machine-readable format called RDF.
Data FAIRification step by step
01
Create interoperable data I
To make data interoperable i.e., machine-readable, we first define a driving user question, e.g., “What effect do different treatments have on the disease severity?”. A driving user question should be defined by the domain expert together with a FAIR data expert. This is to make sure that it reflects both the research interest of the domain expert, but also that it covers the full potentials of Linked Data provided by the FAIR data expert.
02
Create interoperable data II
The question is analysed and the specific data elements and items required to answer the question(s) are identified. Here, this could be: treatment(s) and symptoms/phenotypes that can be linked to disease severity for each patient. Unique persistent identifiers are then assigned to the data. At this stage it’s important to think about what the relations are between the concepts.
03
Create interoperable data III
These will be used to create (or chose) a semantic data model that will be used to ‘guide’ the creation of machine-readable data.
04
Create interoperable data IV
The creation of machine-readable data can be done using the DTL FAIRifier (based on OpenRefine and its RDF plug-in) or the data integration services in MOLGENIS.
05
Create interoperable metadata with clearly defined accessibility and clear description to enable reusability
When we FAIRify rare disease registries we often have no or little metadata to begin with. Therefore metadata needs to be created from scratch. This requires work from e.g., the data owner to describe how the data was created (reusability), a in-house legal expert to define the accessibility, and a FAIR expert to make the metadata interoperable i.e., machine-readable. The creation of machine-readable metadata can be done using the DTL Metadata Editor. We use the Data Catalogue (DCAT) vocabulary to describe metadata, and are currently defining a standard for rare disease registries that defines a way for registries to share and reuse metadata models and address crucial concerns about data access permission and data security.
06
FAIR data point (FDP) I
Metadata is made findable for machines through FDPs and describe under which conditions the data can be accessed and how it can be i.e., reproduced. A FDP needs to be indexed in a search engine in order to be findable. This can e.g., be Google, however, at the Dutch Techcentre for Life Sciences (DTL) efforts are underway to make a ‘FAIR search engine’. The FAIR data point RESTful API uses the DCAT vocabulary and Datacite’s Registry of Research Data Repositories (RE3Data) to provide high level metadata descriptors about data deposits, and to provide instructions to access various distributions of data sets.
07
FAIR data point (FDP) II
Finally, an answer can be found to the driving user question defined in Step 1 by creating sparql queries.
The work on Data Linkage is led by the RD-Connect partners in the Netherlands: Leiden University Medical Center (LUMC) and University Medical Center Groningen (UMCG).