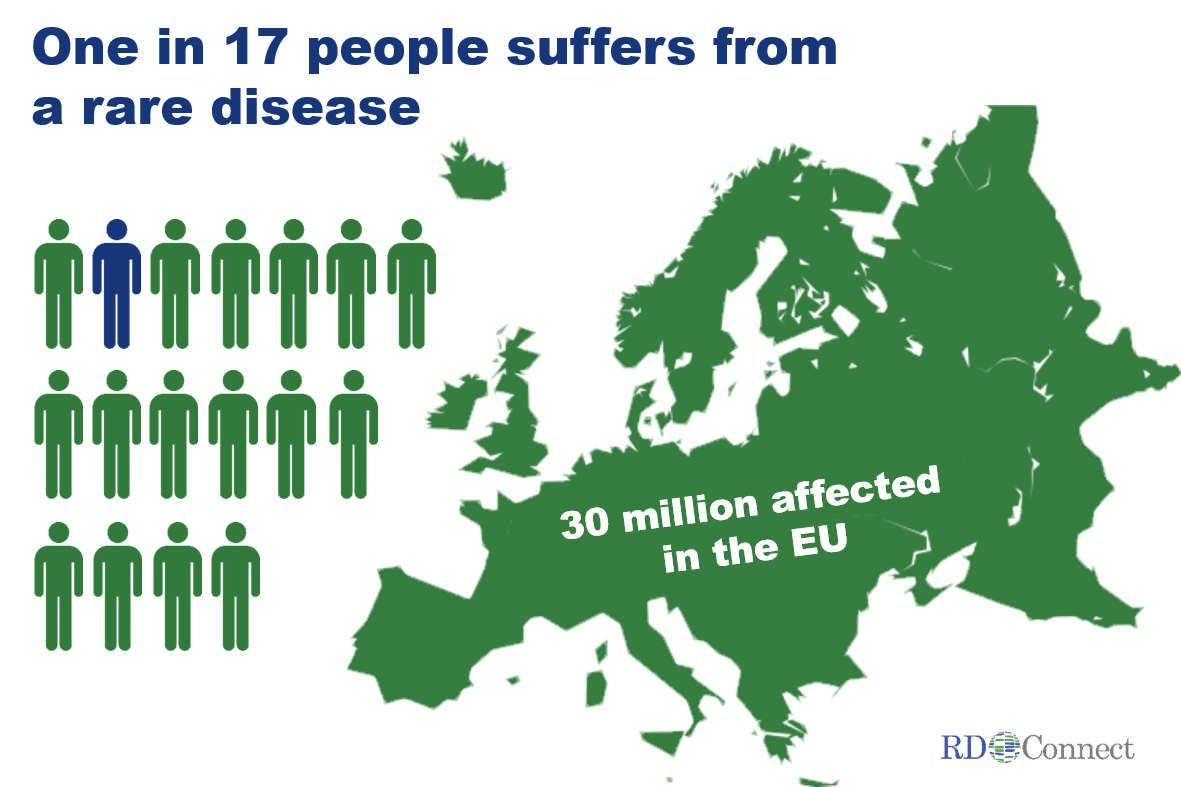
Although individually uncommon, rare diseases collectively affect as many as one in 17 people. In the European Union alone, that is around 30 million people! Around half of them are children. Rare diseases span all areas of medicine and have a substantial impact on public health, society and national economies.
Their rarity and diversity pose specific challenges for research, healthcare provision and for the development and marketing of treatments. Many patients with rare diseases lack timely and accurate diagnosis and even fewer receive tailored treatments.
80% of rare diseases have a genetic component, and the genomics revolution has brought the hope of gene-based treatments for many rare diseases a step closer. The first sequencing of a human genome completed in 2003 required the work of hundreds of scientists for more than 10 years at a cost of over 3 billion €. The same task is now feasible on a single sequencing instrument within days at a cost of less than 10000€, and this cost is continuously decreasing.
Genomics and other newly emerging omics techniques hold the key to new diagnoses and therapies for rare diseases. These new technologies, which allow researchers to look into the way diseases affect the entirety of the DNA, proteins, lipids and metabolites in a human body, are generating data on a huge scale unprecedented in biomedical research. Despite the advances in computing technology, the processing and analysis of data, or even its transfer from one location to another, is difficult and far from routine.
To date, thousands of complete human genomes and hundreds of thousands exomes have been sequenced. Exome sequencing is a technique that captures the part of the genome that holds the “coding instructions” to make the body’s proteins. This has led to an explosion of data that reveals the ways in which one individual’s DNA differs from another’s, and this rapid growth is expected to continue. The limiting factor is now our ability to share and analyse these vast quantities of data, rather than the capacity to generate them. The highest costs are associated with bioinformatics processing of these large datasets. Consequently, new and innovative bioinformatics solutions are required.
However, sequencing is only the first part of the story and doesn’t replace clinical expertise – in fact, combining genetic and clinical data is more important than ever.
The genome sequence of each individual has a several hundred thousand “private” variants that are not found in the general population. The majority of these changes are not directly disease causing, but may still be relevant for gene regulation and the symptom severity. Our current understanding of the underlying biology is often too limited for making appropriate predictions for an individual patient.
Combining and integrating different types of omics: genomics, transcriptomics, proteomics, metabolomics, lipidomics and detailed phenotype data (phenomics) across research centres and across diseases is essential to advance knowledge. While competition between different research groups is a driving force to advance science, harmonisation and sharing of data is ultimately required to compare, combine and make best use of the results. This is especially true in rare diseases, where individuals with the conditions may be scattered across the world.
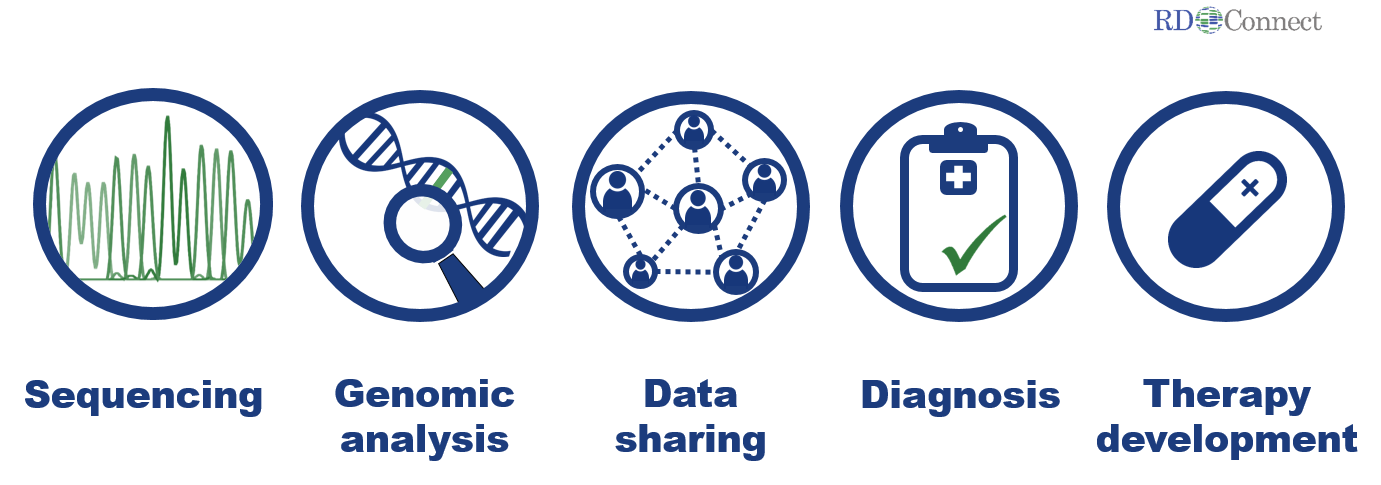
Transnational and trans-disease efforts are thus essential to make optimal use of resources. Patient registries, biobanks and bioinformatics analysis tools are the key infrastructure tools required for omics research. Hundreds of rare disease biobanks and patient registries already exist in Europe alone, and collaborative initiatives in specific disease groups (e.g. Huntington’s disease, cystic fibrosis and neuromuscular diseases) have advanced infrastructure harmonisation in several areas.
A continued bottleneck for research, diagnosis and therapy development is that at present these individual efforts remaining largely “siloed”. Genetic information, biomaterial availability, detailed clinical information (deep phenotyping) and research/trial datasets are hardly ever systematically connected.
To improve diagnosis and therapy, rare disease research needs to be directed towards analysing omics data, linking them with clinical data and biomaterials of individual patients and well-defined patient cohorts. The joint efforts of the three projects – NeurOmics, EURenOmics and RD-Connect – greatly advanced this cutting-edge approach. This may be the first step to revolutionise rare disease research, diagnosis and therapy development around the world.
Read more:
- Sharing data between researchers too often an afterthought in rare disease work – interview with Prof. Hanns Lochmüller (Horizon Magazine)
- Data Sharing in Rare Disease Research (overview on RD-Connect in Rare Revolution Magazine)
- Omics, sweet omics – curing the incurable, one disease at a time (Horizon Magazine)
- RD-Connect, NeurOmics and EURenOmics: collaborative European initiative for rare diseases (European Journal of Human Genetics)