| | | |
Focus on the RD-Connect genomics platform |
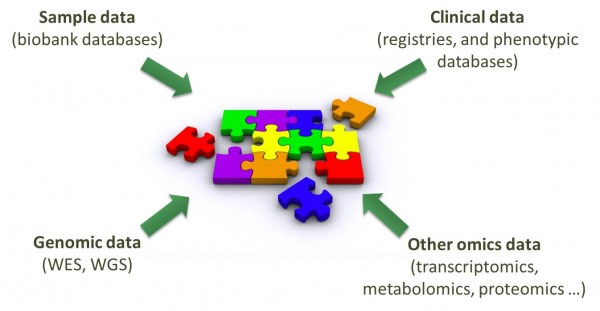
The RD-Connect platform connects phenotypic and omics data for rare disease research As the use of omics technologies expands, it is becoming increasingly essential both to create mechanisms for sharing and analysis of the large amounts of data produced in omics science projects and to develop robust methods of cross-linking these different data types. In particular, it is vital to ensure the linkage of omics data with relevant phenotypic information. Improving the interoperability of different systems and ensuring the systematic connection of detailed clinical information (deep phenotyping) with genetic information, biomaterial availability and research/trial datasets is a requirement in order to deliver concrete benefits to patients in terms of diagnosis and therapy development. RD-Connect is developing an integrated platform in which omics data is being combined with clinical phenotype information and biomaterial availability, accessible online and queryable with a suite of analysis tools. Unlike other omics repositories, which act primarily as filestores, the RD-Connect platform has a user-friendly secure online interface to enable registered rare disease investigators to analyse their data online. EURenOmics (focusing on rare kidney disorders) and NeurOmics (focusing on rare neuromuscular and neurodegenerative disorders) are two flagship omics research projects that feed omics data into the RD-Connect platform. The databases and analysis tools developed in RD-Connect are being piloted in collaboration with investigators from these projects and will be made broadly available in open-source release to the wider community for researchers to uncover more about the genetic causes and pathophysiology of their rare diseases of interest. The RD-Connect genomics platform was officially launched at this month’s ESHG Conference in Glasgow, Scotland, by Steve Laurie (CNAG, Barcelona, Spain) and is currently open for beta-testing by RD-Connect partner project NeurOmics, who are able to analyse their own data through the online interface. Other users will be invited to register later this year. This newsletter will focus on the RD-Connect genomics platform and will provide further information about how you can become a user. |
Data in the RD-Connect platform |
Technology behind the RD-Connect platform The RD-Connect platform has been constructed using state-of-the-art technology adapted to handling the huge amounts of data that will be incorporated over the next few years. The core file system is an Apache Hadoop Distributed File System (HDFS), as used by Facebook and Yahoo, amongst others. This provides many advantages over traditional relational database focussed systems. Firstly, it is fault-tolerant, meaning that if there is a failure within one part of the system, this does not result in the platform failing. Secondly, the distributed nature of data storage allows rapid parallel-processing, providing extremely quick access and recall of requested information. Further, being fully-scalable, as the platform receives more data over the coming years, it will only be necessary to plug-in more hardware - there will not need to be any change to the underlying code. Data is read into the HDFS and processed using Spark before being loaded into Elastic Search (used by CERN and Wikipedia). Storage in Elastic Search allows the platform to make real-time queries on millions of rows of complex data and return appropriately filtered results to the user-interface within a second or two. In order to allow software developers within RD-Connect to integrate their own data and tools into the platform, http APIs are being implemented. This ease of integration of expert tools for advanced analyses is one of the major sources of added-value for users of the RD-Connect platform. The platform has its own dedicated cluster, located at CNAG in Barcelona, and fine-grained controlled access to the data is provided by way of a Central Authentication Service server, managed by CNIO. Technology development will be an ongoing process throughout the lifetime of the RD-Connect project, and as technologies improve, so too will the platform. ~ Steve Laurie and Sergi Beltran, Centro Nacional de Análisis Genómico (CNAG), Barcelona, Spain |
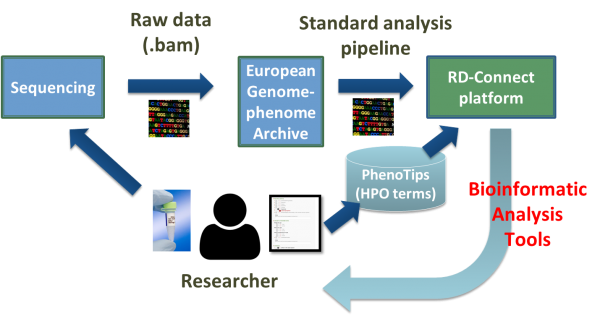
The genomics platform data flow The standard procedure for submitting data to RD-Connect starts with deposition of the raw data in perpetuity in a secure-access repository hosted by the European Bioinformatics Institute. Raw genomic data from collaborating projects is securely deposited in the European Genome-phenome Archive (EGA) and is then realigned and reprocessed through a standard pipeline to ensure cross compatibility of data from multiple projects. The processed data is then held in the central RD-Connect database, where it will be combined with other omics data types plus phenotypic and biomaterial information. Approved researchers can then access data through a data coordination centre that enables comparison of datasets across projects and analysis with sophisticated bioinformatics tools. |
Figure legend: Data flow to RD-Connect |
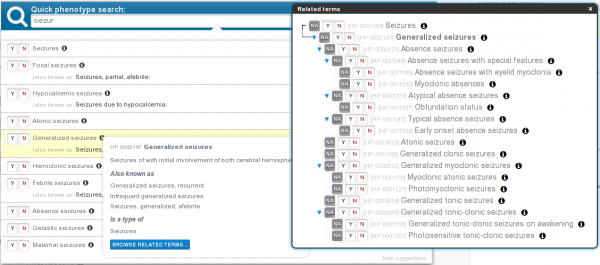
Standardizing clinical data collection using HPO and PhenoTips A key feature of the RD-Connect platform is its integration of detailed phenotypic data about each patient in the system. This makes it possible to query the system for a specific phenotype of interest and link it on an individual patient level with the genomic data held on that patient, and conversely to identify variants of interest in an exome and link back to the phenotype to find out whether the phenotype fits. This is made possible because all projects contributing genomic data have committed to collecting phenotype data in a standardized fashion using the Human Phenotype Ontology (HPO). In recent years the HPO has emerged as a leading system for standardized collection of clinical information about patients with rare diseases, supported by the Global Alliance for Genomics and Health and the International Rare Disease Research Consortium. Many research and clinical projects in the field including the Decipher database and the Deciphering Developmental Disorders project at Sanger, the NIH Undiagnosed Diseases Program, GWAS Central, and the FORGE and Care for Rare projects also use the HPO for all their clinical data entry. The RD-Connect central platform makes use of a user-friendly HPO-based software solution called PhenoTips, which makes it much easier for clinicians to enter phenotypes using the HPO. PhenoTips can be used for any disease, but it can also be customized for specific purposes to provide additional guidance for those entering the phenotypes, and this was used to good effect in a collaboration with RD-Connect’s partner project NeurOmics, where the PhenoTips development team created standardized online forms for clinicians to enter disease-specific phenotypic information for neuromuscular and neurodegenerative diseases on top of the standard PhenoTips instance. But collecting data using the HPO is of value independently of the front-end interface: another partner project, EURenOmics, developed a central HPO-based "phenome" database to collate clinical information from their cohort of kidney disease patients, and thanks to the underlying HPO codes this is also completely compatible with the system. As part of the collaboration with HPO developers (Peter Robinson’s team at Charite in Berlin), terminology workshops regularly take place with expert clinicians in order to augment the HPO, which is still under active expansion, with further phenotypic classes for other rare diseases. |
Figure legend: PhenoTips is a software tool for collecting and analyzing phenotypic information for patients with genetic disorders The value of an ontology like the HPO lies not only in terminology standardization. Since the terms in the HPO are represented in a hierarchical tree structure, if one clinician enters a very precise and narrow phenotype while another enters a more general broad phenotype when seeing the same feature in their patient, the system can still understand that the two phenotypes are linked/similar because the narrower phenotype is represented computationally as a subcategory of the broader one. This is very powerful for the "matchmaking" approaches that are becoming increasingly important as the use of next generation techniques increases, providing a mechanism to link up the many unsolved cases for which final diagnosis cannot be made due to lack of a "second family" or confirmatory case. RD-Connect is participating in the Global Alliance’s Matchmaker Exchange initiative working towards a federated platform (Exchange) to facilitate the matching of cases with similar phenotypic and genotypic profiles (matchmaking) through standardized application programming interfaces (APIs). ~ Rachel Thompson, RD-Connect, Newcastle University |
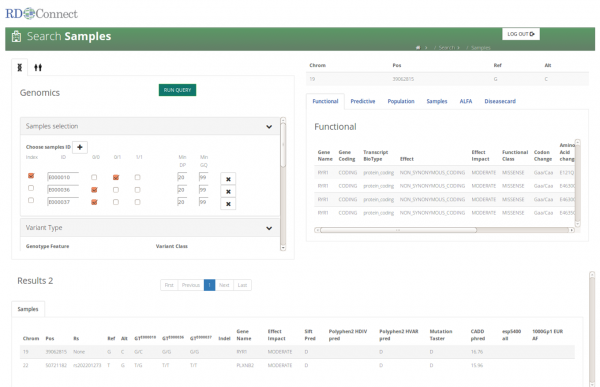
Bioinformatics tools The RD-Connect platform will enable a range of bioinformatics tools to be utilized on data held within the system - both tools that are being further developed within RD-Connect and related projects and external tools that can be linked in through common APIs (Application Programming Interface) and web services. These include variant interpretation and pathogenicity prediction systems, variant/phenotypic "matchmaking" tools, and integrative analysis tools using semantic web applications and frameworks, for improved data integration and access to knowledge. A dedicated work-package, led by Christophe Béroud at Aix-Marseille University, focuses on the development of bioinformatic tools for the platform. The development of such tools will aid data interpretation, data mining and knowledge discovery and help facilitate the discovery of new genes, pathways and therapeutic targets. This work package has already produced several systems that are now available for the scientific community. To date, DiseaseCard, Alamut Functional Annotation (ALFA) and gene-disease relationships in nano-publication format have been integrated. Current focus is on the integration of Exomiser (including PhenIX) to prioritize variants through genotype-phenotype queries, the provision of reliable allele frequencies, the lighting of a Global Alliance for Genomics and Health (GA4GH) beacon and patient matchmaking. Further information and details about the range of bioinformatics tools which are being developed within or alongside the RD-Connect can be found in May’s newsletter or on the RD-Connect website. |
How can YOU interact with the RD-Connect platform? The platform currently contains genotypic and phenotypic data for 367 samples from NeurOmics, and it is anticipated that this number will increase to over 1000 samples by the end of the year, followed by several thousand more in 2016. We will also be incorporating transcriptomic and other -omics data as well. The platform gains power as the number of samples incorporated increases, so if you are working on a rare disease and would like to gain access to the platform, we would be happy to incorporate your samples. In order to do this there are three things you need to do: • Check your consent forms to make sure they allow data sharing for research purposes • Ensure you have a detailed phenotype for each participant • Ensure you have access to the BAM / FastQ files from your sequencing experiments. Further information about the RD-Connect platform including contact details can be found on the RD-Connect website We look forward to hearing from you! |
Publications Factors influencing success of clinical genome sequencing across a broad spectrum of disorders Taylor JC, et al., (2015). Nature Genetics 47, 717–726 In this article published in Nature Genetics, a large group of researchers, including EURenOmics Tubulopathies work package leader Prof. Olivier Devuyst (University of Zurich), present their findings from theWGS500 program which aims to evaluate the clinical utility of whole genome sequencing across a number of human diseases. A total of 217 individuals from 156 independent cases or families across a broad spectrum of disorders in whom previous screening had identified no pathogenic variants were sequenced. The authors found that jointly calling variants across samples, filtering against both local and external databases, deploying multiple annotation tools and using familial transmission above biological plausibility contributed to accuracy. Overall, whole-genome sequencing identified a pathogenic variant in 33 of 156 cases (21%), including 23 of 68 (33.8%) mendelian cases, with the proportion increasing to 57% (8/14) in cases where de novo or recessive models of inheritance were suspected and both parents were sequenced. These results demonstrate the value of genome sequencing for routine clinical diagnosis but also highlight many outstanding challenges. Within RD-Connect there is a dedicated task headed by Peter-Bram 't Hoen and Marjolein Kriek (Leiden University Medical Center) which looks at the impact and challenges of next generation sequencing on clinical diagnostic practice and its implementation into routine healthcare. As more pathogenic variants are identified, genetics centres will have to design new strategies, including new methods for genetic counselling and cross-border referrals, to cope with increasing demands. RD-Connect will monitor these developments. Compromised autophagy and neurodegenerative diseases Menzies FM, Fleming A & Rubinsztein DC (2015). Nature Review Neuroscience 20, 345-57 Autophagy is a basic catabolic mechanism that involves the degradation of cellular components through the actions of lysosomes. Most neurodegenerative diseases in humans are associated with the intracytoplasmic deposition of aggregate-prone proteins in neurons and with mitochondrial dysfunction. Autophagy is therefore an important process for removing such proteins and protecting against neurodegenerative disease. In this review, which is co-authored by David Rubinsztein, NeurOmics lead for the modifier gene identification, prioritization and study work package, the progress that has been made in our understanding of how perturbations in autophagy are linked with neurodegenerative diseases and the potential therapeutic strategies resulting from the modulation of this process is summarised. Intracytoplasmic protein misfolding and aggregation are features of many late-onset neurodegenerative diseases including Huntington disease which is one of the diseases groups being investigated by NeurOmics. Further understanding of the basic biology of autophagy will hopefully provide additional insights into how disease-causing protein variants may affect this degradation pathway. This in turn could lead to the discovery of novel therapeutic strategies and the development of relevant biomarkers, both of which are also aims of the NeurOmics project. A SNP in the HTT promoter alters NF-κB binding and is a bidirectional genetic modifier of Huntington disease. Bečanović K et al., (2015). Nature Neuroscience 18, 807-816 Huntington disease is an autosomal dominant neurodegenerative disease caused by expansion in the polyglutamine-encoding CAG repeat in exon 1 of the huntingtin gene (HTT). Wild-type HTT has a crucial role in the development of the nervous system and is protective against various forms of cytotoxicity, including neurotoxicity induced by mutant huntingtin (mHTT). The expression levels of both wild-type HTT and mHTT have been shown to modify the disease phenotype in Huntington Disease models. In this publication, the authors present in vivo evidence that cis-regulatory variants in the HTT promoter are bidirectional modifiers for age of onset of Huntington Disease. A panel of HTT promoter reporter constructs originating from HD patients identified a SNP (rs13102260:G > A) in a NF-κB binding site that modulated the binding of NF-κB and reduced transcriptional activity of the HTT promoter in reporter gene assays. These results have implications for therapeutic strategies aimed at silencing of the HTT gene in Huntington Disease patients. Genotyping of rs13102260 may also provide relevant prognostic information for a subset of Huntington Disease gene carriers. Furthermore, continued identification of cis- and trans regulatory elements will provide insights into their effect on disease expressivity and identify new targets for disease-modifying therapeutic interventions in Huntington Disease. Identification of genetic variations that influence onset and progression in Huntington Disease, which is headed by co-author Professor Sarah Tabrizi (University College, London), is a key research area for NeurOmics. Also of interest in this paper is the use of cohorts of affected individuals from the European Huntington’s Disease Network Registry, a partner registry of RD-Connect, showing the value of large-scale data collection in registries and also the need to be able to go back to biosamples and patients for further analysis. Full text available here: Taylor JC, et al., (2015).Factors influencing success of clinical genome sequencing across a broad spectrum of disorders. Nature Genetics 47, 717–726 Menzies FM, Fleming A, Rubinsztein DC (2015).Compromised autophagy and neurodegenerative diseases. Nature Reviews Neuroscience 20, 345-57 Bečanović K, Nørremølle A, Neal SJ, Kay C, Collins JA, Arenillas D, Lilja T, Gaudenzi G, Manoharan S, Doty CN, Beck J, Lahiri N, Portales-Casamar E, Warby SC, Connolly C, De Souza RA; REGISTRY Investigators of the European Huntington's Disease Network, Tabrizi SJ, Hermanson O, Langbehn DR, Hayden MR, Wasserman WW, Leavitt BR (2015).A SNP in the HTT promoter alters NF-κB binding and is a bidirectional genetic modifier of Huntington disease. Nature Neuroscience 18, 807-816 |
Workshop in Rome on integration of registry and biobank data The RD-Connect Registry framework and ID-Card Catalogue meeting will take place in Rome, Italy between 30th June and 2nd July 2015. The meeting is co-organised by the Istituto Superiore di Sanità, Leiden University Medical Center and the ELIXIR Research Infrastructure. The meeting is an important opportunity for data linkage experts collaborating with Elixir, registry owners and members of the Core Implementation Group, including: the Ring 14 registry; LeukoFrance database and biobank; the EHDN registry; the UK FSHD Patient Registry, the Global FKRP Registry; the Biobank of the Institute of Rare Diseases Research/Institute of Health Carlos III and the US network of rare disease registries Patient crossroads – to interact and discuss the RD-Connect ID-Card Catalogue of Registries and their integration with the Sample Catalogue. Day one will focus on the interoperability of software for rare disease systems. This will include: (i) the ID-Card and disease matrix of registries; (ii) the new functionalities achieved by linking the ID-Card with Orphanet and (iii) the automatic update of ID-Cards through APIs and the need to adopt a common API format. Participants will also discuss the meaning of different terms such as Common Data Elements (CDEs), Globally Unique Identifier (GUID), Linked Data and Driving User Questions. Registry owners will also be helped by data linkage expert to convert their Case Report Form to linkable, computer-readable data. Day two will focus on clarifying the process flow of how a question is answered by rare disease systems. This will build on the use case questions developed during the 2015 RD-Connect annual meeting to comment on how the integration of phenotype data and biomaterial availability is currently made by researchers and how the integration should be achieved in the RD-Connect platform. Further updates will be given on the patient unique identifier, the RD-Connect Common Data Elements (CDEs) and on the integration of the ID-Card catalogue of registries/biobanks and the sample catalogue developed by Molgenis. The discussion of possible scenarios for the integration of registries and biobanks inside the RD-Connect platform will require clarification of the role of APIs, Linked Data and Ontologies, Globally Unique Identifiers, Human Phenotype Ontology (HPO), CDEs, software services such as COEUS and ID-Card, and the role of different people in the development of these elements. Engineers will also be on hand to discuss the design of an infrastructure that can support federated rare disease resources and the RD-Connect platform. Day three will be a technical meeting for the developers engaged in all of the integration work to interact and discuss strategies and next steps. ~ Sabina Gainotti, ISS and Marco Roos, LUMC |
Launch of Rare Diseases International, the Global Voice for Rare Disease Patients Over 60 patient representatives from 30 countries gathered for the official launch and inaugural meeting of Rare Diseases International (RDI), held at the recent EURORDIS Membership Meeting in Madrid, and to adopt the principles of a Joint Declaration aimed at advocating for rare diseases to be an international public health priority. RDI represents patients and families of all nationalities across all rare diseases and brings together umbrella patient organisations as well as international rare disease-specific federations from around the world. To date, 20 such groups have formally signed up to be a member of RDI and another 50 are expected to join the initiative before the end of the year. Yann Le Cam, EURORDIS Chief Executive Officer, said at the launch, “The foundation of RDI is a historic moment, turning the rare disease patient movement into an international one. By coming together we are creating a critical mass that cannot be ignored. Joining together makes each of us stronger locally and together globally.” He emphasised that globalisation is not a challenge for the rare disease community but part of the solution, commenting, “The complexity of the rare disease community can be united through RDI. Rare diseases are currently ignored on the international agenda. There is a long way to go, but we must look at our diversity as a strength, not a problem.” Durhane Wong-Rieger, President and CEO of the Canadian Organization for Rare Disorders (CORD) added, “CORD has benefitted tremendously from its EURORDIS membership and RDI will be an even greater resource, especially for patient groups in countries that are just developing rare disease policies, by allowing them to draw upon best practices, support and a global patient voice.” Peter L. Saltonstall, President and CEO of the US National Organization for Rare Disorders (NORD), said, “NORD is pleased to join Rare Diseases International and to collaborate with leading patient advocacy groups from around the world to help make rare diseases an important global public health priority.” The main objectives of RDI are: • To promote rare diseases as an international public health and research priority by raising public awareness and influencing policy-making; • To represent members and people living with a rare disease in international institutions such as the World Health Organisation and the United Nations Economic and Social Council; and • To enhance the capacities of members to improve the lives of those living with or affected by a rare disease through information exchange, networking, mutual support and joint actions. RDI is a EURORDIS initiative, created in partnership with national alliances. The preliminary phase of the initiative has been steered by EURORDIS and national rare disease alliances from the US (NORD), Canada (CORD), Japan (JPA), China (CORD), India (I-ORD), the Ibero-American pan-regional alliance (ALIBER) and the International Federation for Epidermolysis Bullosa (DEBRA International). The next annual meeting of RDI will take place in May 2016 in Edinburgh alongside the European Conference on Rare Diseases & Orphan Products (ECRD 2016 Edinburgh). ~ Eva Bearryman, Junior Communications Manager, EURORDIS |
3Gb-TEST Final Symposium: Introducing diagnostic applications of ‘3Gb-testing’ in human genetics The final 3Gb-TEST project meeting will take place on 24 August at the Leiden University Medical Center, Leiden. A major goal of this project is to develop a validated roadmap for the implementation of diagnostic genome sequencing in Europe. In this symposium, the milestones which have been achieved in the project will be high-lighted and there will be presentations on: Genome diagnostics, ethical and quality issues to consider and the future of whole genome sequencing. NeurOmics coordinator, Olaf Riess, (Tuebingen, Germany), will The future of whole genome sequencing: the new role of medical genetics in clinical guiding. The preliminary programme is available and registration is open (free of charge). ~ Ellen Thomassen, Project manager, 3Gb-TEST |
Upcoming events For further information on future events please visit the events page on the RD-Connect website. |
The Human Genetics Society of Australasia 2015 Annual Scientific Meeting, Perth, Western Australia, 8-11th August The Human Genetics Society of Australasia (HGSA) invites you to the 2015 Annual Scientific Meeting (ASM) to be held at the Perth Convention and Exhibition Centre, from 8-11 August 2015 in Perth, Western Australia. The HGSA ASM 2015 promises to be an exciting meeting with a strong scientific program on the theme of Rare Diseases and Indigenous Genetics. The HGSA is the foremost scientific body for those working in the field of Human Genetics throughout Australia and New Zealand. Our membership currently stands at over 1000 members in Australasia. The society also supports a number of special interest groups including: • Australasian Association of Clinical Geneticists (AACG) • Australasian Society of Genetic Counsellors (ASGC) • Australasian Society for Inborn Errors of Metabolism (ASIEM) • Australasian Society of Cytogeneticists (ASoC) • Molecular Genetics Society of Australasia (MGSA) Confirmed speakers include RD-Connect partner, Hugh Dawkins and NeurOmics partner, Nigel Laing Further information is available here The 3rd International summer school on rare disease and orphan drug registries Istituto Superiore di Sanità, Rome, Italy, 21-23rd September RD-Connect partners at theIstituto Superiore di Sanità, Rome, Italy will be hostingan International Summer School which will be focused on the specific aims and needs of registries oriented to clinical research, comprising the study of the natural history of diseases, the assessment of treatment effectiveness and post-marketing surveillance of orphan drugs. The School will train participants on the methodologies and resources available for the establishment of a clinical research registry and on the implementation of successful strategies to ensure long time sustainability of the registry, including data sharing and dissemination activities. The Workshop will consist of brief frontal presentations and practical working groups where participants will learn to make their data interoperable with other sources and databases. The working groups will get together registry owners and bio-informatics experts. This event is open to health professionals, researchers, medical specialists, medical students and representatives of patient associations, who are involved or intend to establish a rare disease patient registry. A selection process will apply based on the participant’s background and role with reference to registry activities. Further information is available here: EMBO Workshop - Molecular Mechanisms of muscle growth and wasting in health and disease, 20 September-25 September 2015, Congressi Stefano Franscini, Monte Verità, Ascona, Switzerland This meeting will focus on the molecular mechanisms involved in muscle wasting diseases including cachexia, sarcopenia and muscular dystrophies. Its focus on disease aspects in skeletal muscle, its interactive format and its small size makes this meeting unique. The conference will take place on September 20-25, 2015 at the Conference Centre Monte Verità, Ascona, Switzerland, the venue of choice for Congressi Stefano Franscini, the international conference platform of ETH Zurich. The conference is limited to 120 participants. As we expect more applicants, we highly encourage to submit an abstract, which will help us to select participants and speakers of short talks. Deadline for abstract submission is Friday 12. June, 2015 Further information is available here. 3rd Ottawa International Conference on Neuromuscular Biology, Disease and Therapy, September 24-26, 2015, Ottawa, Canada The CNMD is hosting the 3rd Ottawa International Conference on Neuromuscular Biology, Disease and Therapy on September 24-26, 2015. After two previous successful neuromuscular disease conferences in Ottawa, the 2015 conference promises to offer an outstanding program emphasizing recent breakthroughs in basic and translational research and clinical discoveries in neuromuscular disease. The Conference is structured for both basic researchers and clinicians and will feature internationally-recognized invited speakers highlighting advances in all aspects of NMD research, including novel techniques to diagnose NMD, biology of disease pathogenesis, expanding clinical phenotypes, basic muscle and stem cell biology, and promising therapies to treat these devastating disorders. As in past years, trainees are encouraged to attend and participate – selected abstracts will be featured for platform presentation during the scientific sessions, and all posters are eligible for top poster awards. Confirmed speakers include RD-Connect coordinator Hanns Lochmüller, RD-Connect Associated partner Kym Boycott and NeurOmics infrastructure workpackage leader Volker Straub Further information is available here. TREAT-NMD Alliance Bi-Annual Translational Sciences Conference. December 6 - December 8, Washington D.C., USA This conference will be a fantastic opportunity for patients, academics, clinicians, patient registry curators and industry representatives, to get together to network, learn and exchange ideas about translational research. RD-Connect will be speaking about the role of neuromuscular registries in -omics research and the wider RD-Connect project. Further information is available here |
Why did I get this email? You received this email because you are associated with RD-Connect, EURenOmics or NeurOmics or because you signed up online. We will send out around one email per month with news of relevance to these projects and to IRDiRC. If you don't want to receive any further newsletters, you can unsubscribe using the link below. If you're reading this online or if it was forwarded by a friend, you can sign up to future editions here. |
| | | | | |